Islands of Fertility: Oak Tree vs. Buckwheat Savannah Soils
Between Sandy Creek and Chalone Creek, waypoints ending in -P006 through -P009.
Samples collected under Live Oak (SSGG-spring-05-P006, SSGG-spring-05-P009) correspond to the soil described at pit S04CA069007K (Corralitos). Samples collected under Buckwheat (SSGG-spring-05-P007, SSGG-spring-05-P008) correspond to the soil described at pit S04CA069004K (Toags). Bulk density data were collected via ring method at all 4 locations. Compliant-cavity method data were taken at pits P006 and P007.
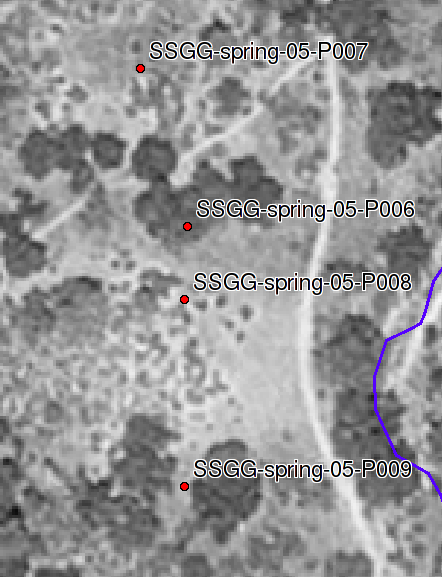
Islands of Fertility: Sample Sites: Sample sites for islands of fertility project. Pits were described and sampled by the Soils and Biogeochemistry Graduate Group, Feb. 2005.
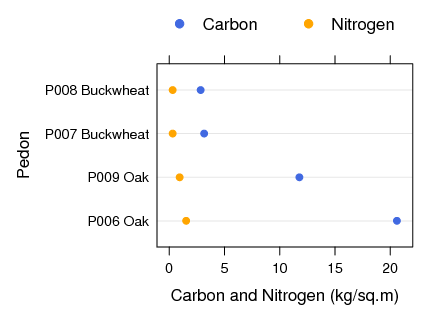
Total C and N Under Oak vs. Buckwheat
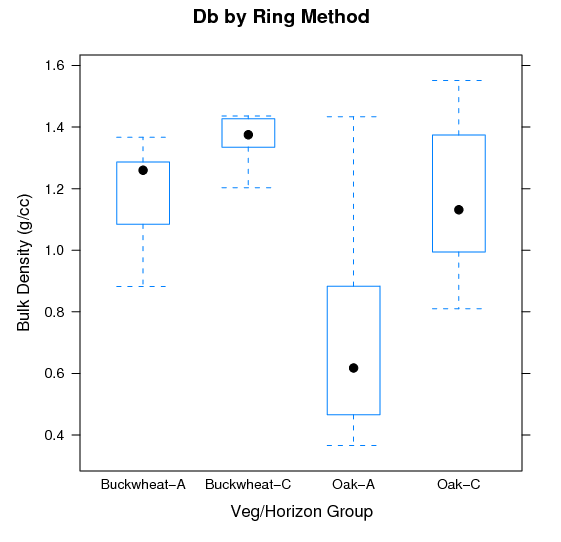
Islands of Fertility: Differences in Bulk Density: Variation in Db as a function of cover type and major horizon designation.
Analysis in R
Projects
- BMP's for Irrigated Agriculture
- Pedology and Soil Survey
- Geographic Nutrient Management Zones for Winegrape Production
- GIS and Digital Soil Survey Projects
- New Technologies in Soil Survey
- Other Information
- Pinnacles National Monument
- Terrain Classification Experiment 2: GRASS, R, and the raster package
- Images from Pinnacles Soil Profile Analysis
- Accessing PINN Soils Data in Google Earth
- Computing terrain-specific slope classes by region
- Finding pockets of soil between the Pinnacles
- Islands of Fertility: Oak Tree vs. Buckwheat Savannah Soils
- Pedon Data collection and entry graphs
- Restored 1933 Geologic Map of Pinnacles
- Soil Color Ideas
- Soil Properties by Parent Material and Rock Type
- Some panoramic pictures
- Insolation Time Experiments
- Pinnacles National Monument
- Rangeland Soil Management and Hydrology